What Is Life?
What is life? This question keeps many people awake at night and has led to rafts of definitions, some involving features (procreation, metabolism and so forth), some involving chemistry, entropy or energy flux and some are of a more philosophical bent. A BBC article noted there are over 100 definitions of life and it claims all are wrong.
And indeed it does appear to be quite a challenge - every hard and fast rule is violated somewhere in biology. There are things that are clearly alive, but never replicate. Plants typically do not move yet are extremely vital. Viruses do make copies of themselves, but need a host to do so. Are they alive? And once you have it all figured out, some wise person comes up with a crystal that somehow manages to create copies of itself, and points out that these copies even ‘inherit’ characteristics. Is the crystal alive? Similarly, “fire” consumes elements, moves and even replicates. Yet calling fire a life form seems odd.
As fascinating as these discussions are, they are also somewhat silly. It is a bit like a bunch of people standing in an electronics store and pointing at phones, watches, cameras and TVs and asking which of these are computers. They may note that the watch exhibits many phone-like features, and a modern phone is clearly a computer, but most watches can’t actually make calls without a phone nearby. They may also argue that the modern TV is not functional without an internet connection, so does it qualify as a computer? They are sure however that an SD card is not a computer, it only stores data.
Whereas to seasoned technologists this is not even a discussion - all of these things are computers of course, and in fact almost all of them by now run a variant of UNIX. The architecture answers the question for us - all of them have billions of transistors that make up CPU cores, RAM, and there is a operating system that runs on top of these. To us, this is not worth lying awake for at night since we know what is going on inside TVs, phones and watches. And in fact, the lowly SD card clearly also is a computer running a whole operating system.
It turns out that all life on Earth also shares a single architecture. And at least on Earth, that massively simplifies the discussion - Anything built on that architecture is life, and we have yet to encounter anything that is remotely living that is not based on that architecture.
Now, some people don’t have the luxury of ignoring anything beyond Earth, which is why NASA’s definition of life is rather fascinating. A fun read on the subject is Defining Life by Steven A. Benner. And we may also wonder if a sentient computer program might one day be called alive of course. As a thought experiment, thinking about that we would consider to be “life” can be very fascinating.
But here on Earth, for now the question of what is life can easily be answered with “whatever runs on the Earth’s single architecture of life”. So what is that single architecture?
RNA, DNA, Proteins
None of what follows is in any way controversial. At the same time, the story is not often told like this, but everything here can be found in any molecular biology textbook
So what is life? How does it actually work? How is it an architecture? How does it get stuff done? We are still exploring the complexities of life, and this will likely occupy us for centuries to come, but the basics are now abundantly clear. The total complexity however is astounding, which is only to be expected for a many billion year old software project.
Software? Let’s start with the basics. All (*) information in life is stored as nucleotides, which are simple molecules. There are 4 different nucleotides and we call them A, C, G and U/T.
“Actually”, a real biologist will now interject, DNA is not the whole story. Every cell comes from another cell. And much like the source code of a compiler is not yet a compiler, because we need to bootstrap it, if all we had was DNA, we would not be able to create a new organism. How much non-DNA information is required for life is not clear.
A typical bacterium runs on a few million nucleotides stored in a long molecule called DNA. Some bacteria have additional smaller DNA molecules called plasmids, which we can literally regard as ‘plugins’. Plasmids can for example confer antibiotic resistance to a bacterium. All in all, it adds up to perhaps 750 kilobytes of data. Tightly curled up, the 1 millimeter long main DNA molecule fits within a micrometer sized bacterium.

Utricularia gibba, record small genome for a vascular plant (Source: Wikipedia)
More complex forms of life have billions of nucleotides, but interestingly don’t specifically need to do so it appears. There is complex plant life based on only a few megabytes of DNA. A human being has around 750 megabytes of DNA, but most of this consists of repetitive stuff - we could likely fuzzily compress our DNA to only dozens of megabytes of information.
Amino acids
There are 20 of these, and they are encoded in DNA using 3 nucleotides which we together call a codon. ‘AAC’ is an example, or ‘CCG’. Since there are four nucleotides, there are 444=64 different codons, but only 20 different amino acids. Many codons code for the same amino acid therefore.
The conversion table between codons and amino acids is, with very very minor differences, universal for all forms of life on Earth.
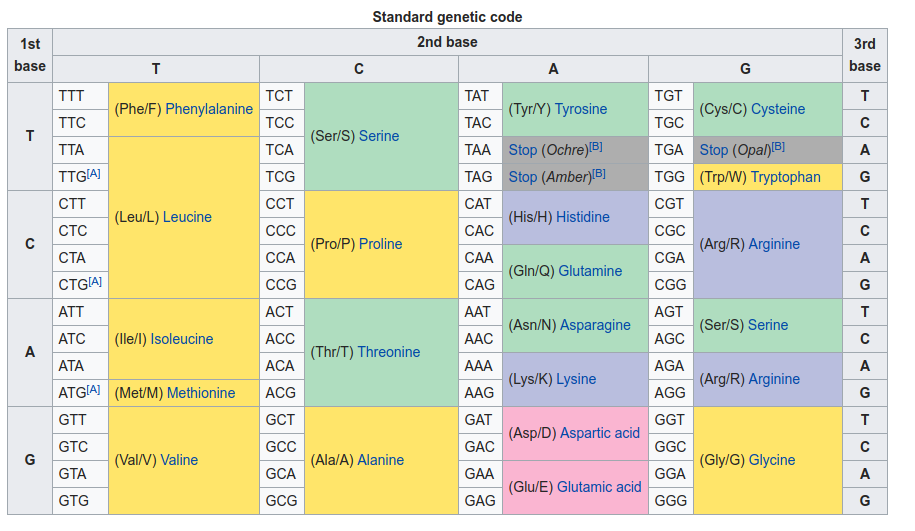
DNA/Codon table (Source: Wikipedia)
Amino acids can be strung together to form proteins. Proteins are small to very large molecules that can speed up or block chemical reactions, exert force or generally perform (or enable) almost any chemical or physical process.
RNA
DNA can be seen as ‘ROM’, as in, the data mostly just lies there. There is however machinery that can take (relatively) small pieces of DNA and turn them into RNA molecules. These eventual RNA molecules contain all or a subset of the information found in DNA.
Cells have separate bits of machinery called ribosomes that turn RNA molecules into chains of amino acids, which we then call proteins.
Proteins
Proteins can do many things, which is impressive given they are built from only 20 components. Some proteins have only an internal meaning - insulin for example signals that cells should change their behaviour, but by itself insulin doesn’t do or build anything.
Other proteins, like for example the excitingly named cryptochromes detect light and even magnetic fields. Yet other proteins sense PH levels, temperature or glucose concentration. They react to such phenomena by changing shape (conformation) after catching sufficient photons or warming up beyond a certain temperature, for example.
Proteins are also enzymes that speed up specific chemical reactions by dozens of orders of magnitude (!), making it fair to say that proteins actually build other molecules.
In addition, proteins can also “just” be structure for a cell or a cell component.
Recap
DNA is large and mostly static. From this DNA, snippets are transcribed into separate RNA molecules. Such RNA molecules may be picked up by ribosomes to be translated into amino acids which are then chained together to form proteins.
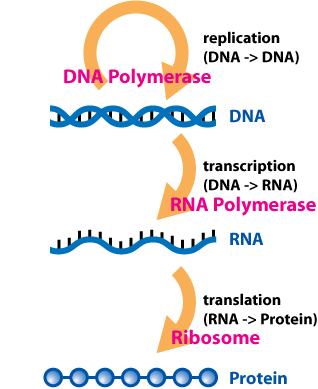
Central dogma of life (Source: Wikipedia)
And proteins are the building blocks of life that also sense and do things.
Interactions
This being nature, almost everything interacts. For example, some proteins get stuck to specific bits of DNA. When stuck there, they can block that part of DNA from being read, so no RNA will be formed, nor will there be any new production of the protein it might have encoded for.
But conversely, if a protein gets attached slightly ahead of a bit of DNA, it can attract the DNA to RNA copying machine and promote the production of RNA and thence other proteins.
Importantly, proteins that change shape (conformation) often only stick to their part of DNA in one of the shapes, but not in the others.
RNA meanwhile can similarly influence or block parts of DNA from being turned into RNA.
Ok, impressive, but show me the software
Up to here we’ve seen elements that might make algorithms possible - proteins can influence the generation of other proteins, proteins can sense other proteins and the environment. This clearly smacks of ‘if’ statements.
The “operating system” of life is on one hand highly recognizable for a software person, but at the same time also downright alien.
Assume we have a DNA molecule and that the machinery to copy DNA to RNA already exists, as do the ribosomes that can turn RNA molecules into proteins.
Initially these things just float around in Brownian motion. Some parts of DNA are shaped so they attract the copying apparatus from time to time under normal circumstances and these parts of DNA are then part of the ‘housekeeping genes’.
Such genes might for example create a protein that is sensitive to temperature, perhaps causing the protein to change shape at (say) 35 degrees C. This protein then functions as a thermometer. Let’s now assume that in its “cold” shape, it binds close to a part of DNA that encodes for a protein that creates heat.
If it becomes too cold, the thermometer protein flexes, and binds to that part of DNA, where it stimulates the conversion of ‘downstream DNA’ into RNA into a protein that (say) stimulates a reaction involving glucose that releases heat, thus warming things up again.
But then what?
The thermometer protein promotes the production of the heat generating protein, and this duly heats up the cell.
As the organism warms up, the thermometer protein will change back conformation and the glucose-reaction-inducing-protein will no longer get generated.
However, once made, that protein will hang around and continue to stimulate the glucose reaction, eventually overheating the cell and likely killing the organism. Not good.
There are many regulatory loops running in every cell, some of incredible sophistication. Nature applies very advanced techniques to precisely regulate PH levels for example.
In our thermostat case, a typical ’nature’ solution is to have the glucose reaction stimulating protein itself be temperature sensitive, causing it to break down quickly at (say) 40 degrees C.
Alternatively, the protein might just unstable at any temperature, and the whole thing only works if it keeps being generated because it is “too cold”.
Beyond on/off
The thermostat described above implements a single ‘if’ statement. As noted earlier, proteins can attach to DNA in such a way that they enhance the ’transcription’ and ’translation’ into new proteins, but they can also inhibit this process.
Different proteins can thus combine to create arbitrarily complex ‘if’ statements. In one of the best studied algorithms, the lac operon in many bacteria, there is a condition to only convert lactose into glucose if 1) a shortage of glucose has been detected 2) lactose is actually present.
The thermostat example above is entirely fictional, but the lac operon makes for fascinating reading on how real natural algorithms work.
Note that since this is nature, the if-statement is more of an ‘if-ish’ statement. Based on the concentration of glucose, for example, more or less might happen - not many things are completely black and white in biology.
In addition, here we’ve mentioned only one mechanism that inhibits or promotes the production of proteins - it turns out there are many more modulation mechanisms all along the way, mechanisms which not only modify the speed of creation but that can for example also conditionally leave parts of genes in and out of proteins.
In more complex forms of life, whole ‘master switches’ are available to inhibit or promote swathes of genes at a time, much like a (musical) organ has registers to enable the generation of specific sounds by changing many stops at once.
The architecture of life is a truly powerful event driven system.
So, what IS life?
The description above is applicable without change for all kinds of bacteria, all plants, all animals and everything in between. There is a vast divide in life between prokaryotes (at least two kinds of bacteria) and eukaryotes (us, animals and plants), with the prokaryotes on the one hand being vastly simpler, but on the other hand also exhibiting far more versatility. Eukaryotes come with significantly more infrastructure and even more ways of modulating the workings of DNA.
But both eukaryotes and prokaryotes share the exact same molecular architecture described above, to the point that bacteria are perfectly willing to ’execute’ human DNA to (for example) make insulin for us. Interestingly enough, this does work out of the box, but if some slight adjustments are made for the bacterial dialect, insulin production speeds up by orders of magnitude. Given the billion years old split between bacterial lineages and us, it is stunning that the only difference is ‘dialect’.
So what is this dialect? As noted there are 64 codons encoding for only 20 amino acids, so most amino acids have several codons that describe them, but not all of these codons are as frequently used in all organisms. Bacteria have a very different ‘codon bias’ compared to our genes, and transcribing DNA using codons common in humans is a lot slower than if these unfamiliar codons are changed for ones common in bacteria.
But what about viruses?
Viruses also use DNA and RNA, but they do not host the rest of the machinery to make proteins. This might confuse the notional philosophers in the electronics store mentioned earlier, but it need not confuse us, because the situation is well known in our world. Viruses are not alive, but they are software that runs on living things.
If the protein that encapsulates the virus software infects a cell, its own DNA (or RNA) will run on the machinery of that living cell. If successful, this turns the cell into a factory for producing new viruses, which when released can then go on to infect yet further cells.
In computing terms this is actually closer to what we’d call a worm.
Summarising
The definition of what life exactly is continues to be fascinating, but at least on Earth, for the life forms we know about, things are pretty simple. If it runs the architecture of life, it is life. If it runs on that architecture hosted by another cell, as for example viruses do, it is not life but software - but it is software that comes alive once inside a host that can run it.
